Decoding the Gut: How AI Mapping Is Redefining Type 2 Diabetes
Date
April 15, 2026
Credits

Date
April 15, 2026
Credits
Medical providers featured in this article

In Brief
- Gut health is a critical upstream driver of complex chronic diseases, including insulin resistance and Type 2 diabetes.
- World leader in interventional gastroenterology BarhamAbu Dayyeh, MD, MPH is rewriting the molecular language of the gut to move his field closer to disease modification.
- By integrating digital pathology, spatial transcriptomics and machine learning, duodenal biopsies can evolve from exclusionary tests into diagnostic tools that enable metabolic—disease subtyping and precision-therapy selection.
- Regenerative electroporation therapy (re-cellularization via electroporation, or ReCET) uses nonthermal pulsed electric fields to induce mucosal regeneration, with early-phase data showing tissue-level normalization and clinically meaningful improvements in glycemic control.
- Together, AI-enabled diagnostics and regenerative endoscopy signal a shift from symptom management toward upstream disease modification, expanding gastroenterology’s role in precision metabolic care.
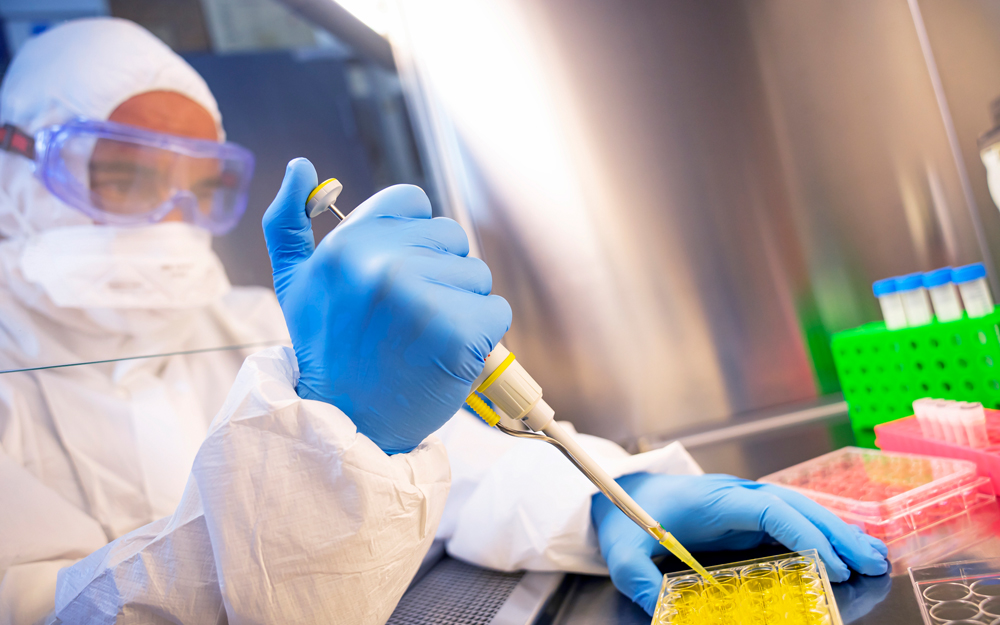
The gastrointestinal tract is a dynamic, information-rich interface between the external environment and human physiology. Gut health is emerging as a critical upstream driver of complex chronic diseases, including insulin resistance and Type 2 diabetes.
Barham Abu Dayyeh, MD, MPH, a world leader in interventional gastroenterology and a prolific clinical investigator, is executive director and associate dean for innovation at Cedars-Sinai and serves as director of both Interventional Gastroenterology and System Integration in Advanced Endoscopy.
“Type 2 diabetes affects 1 in 8 Americans, yet we still define it by blood sugar levels,” he said. “If we want to change the trajectory of this disease, we have to understand—and target—the tissue-level pathophysiology that drives this disorder.”
Abu Dayyeh leads a multidisciplinary program that aims to map diseases like Type 2 diabetes at the level of the duodenum by leveraging AI, spatial transcriptomics and advanced digital pathology. Central to this work is identifying metabolic defects in the duodenum, what he refers to as “duodenopathy”; the structural and molecular abnormalities that underlie insulin resistance, metabolic dysfunction and inflammation.
Discoveries spoke with Abu Dayyeh about his latest research in the field.
Why are you interested in interrogating the gut?
Our work began with a clinical paradox: Certain gastrointestinal interventions produce metabolic improvements that cannot be explained by weight loss alone. Traditional markers such as hemoglobin A1c, fasting glucose and insulin resistance tell us what’s happening downstream, but they don’t explain where the disease originates.
The gut, and especially the duodenum, sits at the crossroads of nutrient sensing, immune signaling and hormonal regulation, so it was a logical place to look. We wanted to know whether disease-specific biology was embedded in the tissue itself—and whether pinpointing that pathophysiology could help us modify disease trajectory.
Type 2 diabetes affects 1 in 8 Americans, yet we still define it by blood sugar levels. If we want to change the trajectory of this disease, we have to understand—and target—the tissue-level pathophysiology that drives it.
How did you build the AI“disease map” of the duodenum?
It was a global effort conducted across Cedars-Sinai, Mayo Clinic and Vanderbilt, as well as international partners—and it began with human patients, not animal models. We obtained duodenal biopsies in carefully phenotyped cohorts with and without metabolic disease and digitized them, essentially creating a 3D, multilayered tissue view.
Using spatial transcriptomics, a molecular technique that measures gene activity in tissue samples, and other advanced technologies, we generated 3D cellular renderings that capture how cell populations and neighborhoods “talk” to eachother. Then we applied machine-learning models to analyze these maps and identify a recurring spatial transcriptomic signature—a molecular QR code—that reliably distinguishes metabolic duodenopathy from normal tissue.
What has AI mapping revealed about Type 2 diabetes?
AI mapping uncovered a reproducible molecular signature in the duodenal mucosa that reflects immune dysregulation, epithelial dysfunction and altered intercellular communication networks. Importantly, these molecular patterns correlate with insulin resistance and glycemic severity, suggesting the tissue itself reflects—and likely contributes to—systemic metabolic dysfunction and inflammation.
How could these findings translate to patient care?
The duodenum is readily accessible during routine upper endoscopy, and tissue sampling is already part of standard GI care. If biopsy acquisition and computational analysis can be standardized, duodenal tissue could allow clinicians to subtype metabolic disease, predict progression and select therapies based on underlying pathophysiology.
Just as laser resurfacing stimulates renewal when skin ages, we are testing a similar treatment for Type 2 diabetes. Re-cellularization via electroporation—ReCET—delivers calibrated electrical pulses endoscopically to trigger duodenal regeneration, recalibrating abnormal metabolic and inflammatory signaling without surgery, implants or permanent anatomical change.
How does this approach compare mechanistically with GLP-1 receptor agonists, and can these strategies be combined?
GLP-1 therapies areeffective for many patients, but our spatial molecular analyses suggest they do not reverse the underlying duodenal molecular and inflammatory dysfunction in Type 2 diabetes. That does not diminish the therapies’ role; it highlights that they act through different biological mechanisms. Looking ahead, AI-guided diagnostics may help determine which patients would benefit from pharmacologic therapy alone, who would benefit from regenerative therapy and who might require a combination approach.
What excites you most about this area of research?
Most current treatments for metabolic dysfunction focus on managing hyperglycemia rather than reversing disease pathophysiology. If your symptom is high blood sugar, we give you medications and insulin to lower your blood sugar. But diabetes often continues to progress and escalate, leading to kidney disease, cardiovascular events and premature mortality.
This work introduces the possibility of intercepting disease earlier and targeting the pathophysiologythat drives progression. If we can read—and then rewrite—the molecular language of the gut, we will move closer to disease modification, not just symptom control.